Pooled Lentiviral shRNA Screens to Identify Essential Cancer Pathways
Screening with RNAi is a powerful method for perturbing gene activity in cultured cells to enable gene function, pathway analysis, and target identification. shRNA is utilized for stable gene knockdown and two formats are commonly used for shRNA screening: arrayed libraries and pooled libraries.
The arrayed screening format has been the primary format for identification of novel gene targets. Arrayed screening involves single shRNA clones in each individual well of a plate, which allows for the simple detection of positive hits. While simple and convenient, arrayed screens require hundreds of plates, the use of expensive equipment such as liquid handlers, and are cost prohibitive to most users. In addition, arrayed screening is not applicable for in vivo use. A pooled screening approach involves multiple shRNA particles targeting many genes in one pool which is used to transduce a plate of cells. With pooled shRNA screening, a deconvolution strategy must be utilized to determine which phenotype is a result of a specific shRNA integration event. A pooled approach is more affordable, can be performed by one researcher within a week, and does not require any specialized equipment.
Possemato et al. detailed an in vivo pooled shRNA screening approach to identify that the serine synthesis pathway is essential in breast cancer1. A list of 2,752 genes gathered from maps of metabolic pathways and cross-referenced to the KEGG Database were compiled and screened. Genes were scored based upon three properties; 1) up-regulation in tumors compared to normal tissues, 2) high expression level in breast cancer, and 3) association with the stem cell state. Genes scoring in at least two of these categories, as well as those at the top of each category, were selected to define a high priority set of 133 metabolic enzyme and transporter genes. Lentiviral shRNA vectors targeting the selected genes, with a median 5 shRNAs per gene, were assembled and used to generate two lentiviral shRNA libraries. One library contained 235 distinct shRNAs targeting transporters and control genes and the other contained 516 distinct shRNAs targeting metabolic enzymes and control genes. Human MCF10DCIS.com cells were then used to screen for depletion of shRNA post tumor formation in mouse mammary fat pads. This resulted in a list of 16 targets, several of which had previously been identified as having important roles in cancer.
One of the genes that overlapped between screens was phosphoglycerate dehydrogenase (PHGDH). PHGDH is commonly amplified in estrogen receptor negative breast cancers and melanomas. It was shown that PHGDH drives serine synthesis and its knockdown via shRNA results in reduction in serine levels, inhibition of cell proliferation in vitro, and reduction of tumor size in vivo, suggesting a requirement for this pathway in this tumor type. This was the first in vivo pooled shRNA screen using The RNAi Consortium (TRC) library to identify potential therapeutic targets for breast cancer. Several targets of potential clinical significance were identified.
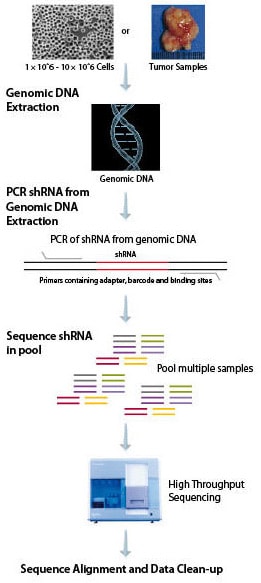
Because there are multiple shRNAs in each pool, developing an appropriate method of identifying the particular shRNA having an impact in a pooled screen is extremely important. We have developed a comprehensive deconvolution solution utilizing deep sequencing with proprietary PCR primers to amplify each shRNA. One of the many advantages to utilizing this method is the ability of deep sequencing to clearly identify each shRNA by sequence and to quantify each shRNA. Deep sequencing will provide digital counts on the number of times each shRNA is identified in each sample, affording no limit on the dynamic range of detection. This is superior to hybridization-based detection, which may lead to DNA saturation. When the oligo becomes saturated, the ability to quantify individual shRNAs is lost. Saturation can also result in false positives due to non-specific DNA binding of similar sequences. Deep sequencing can be applied to both custom and whole-genome samples, which increases the flexibility of deconvolution. MISSION® shRNA is fully sequenced during deconvolution, allowing for the utilization of a barcode to identify different samples during sequencing. By multiplexing multiple samples into a single lane, we are able to decrease costs and increase efficiency, enabling researchers to save time and money while advancing their drug discovery.
DNA Preparation and Deconvolution Workflow