Fragmentation of Chromatin for ChIP Applications
Fragmentation ensures that protein/DNA complexes in high–molecular-weight chromatin are soluble and accessible to your ChIP antibody. The method of fragmentation that you choose should depend on whether you performed N-ChIP or X-ChIP. If you are performing N-ChIP, then you should fragment the DNA with appropriate enzymes (such as micrococcal nuclease), because mechanical shearing methods will disrupt native histone/DNA associations. If you are performing X-ChIP, you may choose either mechanical sonication or enzymatic digestion to fragment your DNA. In either case it is important to optimize shearing conditions and use those exact same conditions for each experiment to reduce the potential for variability in the starting chromatin (Figure 1).
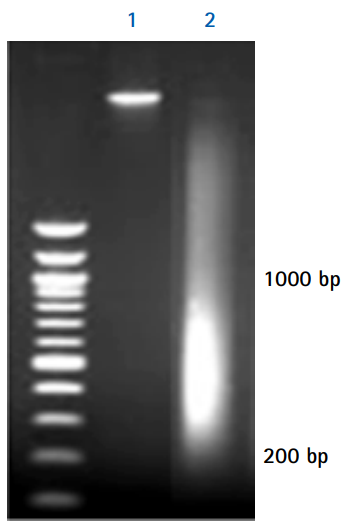
Figure 1.Agarose gel electrophoresis analysis of purified DNA fragments. The purified DNA fragments have undergone sonication (Lane 2) or no sonication (Lane 1), proteinase digestion, crosslink reversal, extraction and precipitation. We recommend that you analyze the DNA on an agarose gel after each sonication experiment.
Chromatin Fragmentation Tips
- It may be challenging to replicate published shearing protocols without optimization. This can be especially true if your instruments differ from those used in those protocols. There are a variety of sonication instruments, both water bath and probe-based. When using a probe-based sonicator, select a tip that is appropriate for your sample volume. In any event, shearing parameters should be optimized for your sample volume, cell density and cell type.
- Optimization should include the power settings (time on versus time off/rest period) and the number of shearing cycles required to obtain DNA fragments between 200-1000 bp in length. This size is important since downstream analysis assays are typically designed to detect only 100-200 bp amplicons containing a binding site of interest. To perform successful optimization, vary only one parameter for each optimization experiment; for example, keep the power setting constant while varying the number of cycles.
- The efficiency of sonication also varies with each cell type and the number of cell equivalents. You may need to perform separate optimizations for different types of cells or tissues.
- Be careful of time and power settings. Over sonication and too high of a power setting can damage the epitopes you are trying to preserve for the immunoprecipitation step. This will reduce your ChIP signal.
- Always keep lysates ice cold and sonicate at intervals rather than continuously, as sonication produces heat, which can denature chromatin. Avoid foaming during sonication. Foaming can result in surface denaturation of proteins and can result in the loss of chromatin in air bubbles. To avoid this, begin with low power settings and gradually increase the power.
- When optimizing conditions, analyze the length of DNA fragments by agarose gel electrophoresis after each sonication cycle. For optimal sizing, you should perform additional cleanup steps to purify the DNA before electrophoresis. Large, insoluble protein: DNA:RNA complexes resulting from inadequate shearing, may clog the pores of the agarose gel and retard the electrophoresis process. Purify the DNA by digesting the proteins, reversing crosslinks, performing phenol:chloroform extraction and precipitating the clarified DNA.
NOTE: that heterochromatin may be resistant to sonication and this may reduce the yield of chromatin.