Protocol for Purification, Optional Reduction, and SEC-MS Analysis of a Monoclonal Antibody
Section Overview
- Workflow for Intact and Middle-Up Mass Analysis of Adalimumab
- Characterization of Monoclonal Antibodies and the Role of SEC-MS in Intact and Subunit Mass Analysis
- General Procedures - Sample and Reference Preparation and System Setup for Antibody Purification and SEC-MS
- Adalimumab SEC-MS Intact and Middle Up Analysis Results
- Conclusion
- Related Products
Workflow for Intact and Middle-Up Mass Analysis of Adalimumab
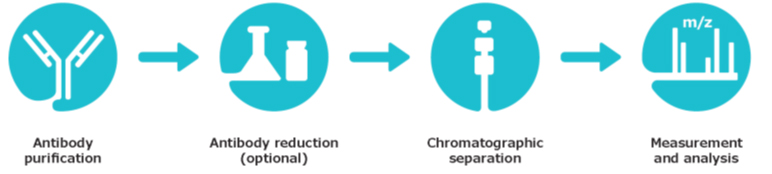
A complete SEC-MS workflow has been developed to simplify intact mass analysis of both non-reduced and reduced monoclonal antibodies (mAbs).
In detail, it includes:
- Antibody purification using immobilized protein A
- Antibody reduction procedure (optional)
- Mass spectrometer calibration
- System suitability test utilizing a recombinant human monoclonal antibody reference
- One generic SEC-MS method suitable for sample separation and analysis of both non-reduced and reduced monoclonal antibodies
Characterization of Monoclonal Antibodies and the role of SEC-MS in Intact and Subunit Mass Analysis
Monoclonal antibodies (or immunoglobulins - IgGs) are large glycoproteins with a molecular weight of approximately 150 kDa (150,000 g/Mol). They are composed of two identical light chains (LC, molecular weight ca. 25 kDa each) and two identical heavy chains (HC, molecular weight ca. 50 kDa each) linked through covalent inter- and intra-chain disulfide bonds. They are utilized for the treatment of various types of cancer, and other diseases such as multiple sclerosis, Alzheimer’s disease, and migraine.
Careful and thorough characterization of therapeutic mAbs is essential for ensuring drug safety and efficacy. mAbs are typically manufactured in mammalian host cell lines in bioreactors, generating a large number of heterogeneous drug molecules. It is important to establish critical quality attributes (CQAs) for each mAb and demonstrating that production batches are within acceptable limits are requirements for both innovator and biosimilar therapeutics.1,2
In many cases, the characterization of an antibody-based drug is performed using a specific chromatographic technique (e.g., size exclusion, reversed phase or hydrophilic interaction liquid chromatography, respectively – SEC, RP or HILIC)3,4 coupled with mass spectrometry (MS). This combination allows for different types of analyses to be carried out, e.g., accurate mass measurement of the intact mAb and subunits, peptide mapping, and the determination of post-translation modifications such as glycosylation, oxidation, and deamidation.
Several techniques are applied to simplify antibody analysis by either fragmentation or removal of glycans. The latter can be performed by treatment with PNGase F, whereas proteolysis with IdeS5 or reduction of inter-chain disulfide bonds with reducing agents, such as dithiothreitol, result in the formation of different antibody fragments with masses of 25 – 50 kDa. Various combinations of these techniques can be applied. For analysis of mAbs in cell culture supernatants, these may be combined with a preceding affinity purification step.6 These approaches are referred to as intact mass and middle-up analysis methods.7 The former term relates to the measurement of the mass of an intact mAb without controlled dissociation being performed. Such an experiment reveals information about stoichiometry, proteoforms, and modifications. Middle up experiments include mass measurement after cleaving mAbs into several large fragments/subunits via chemical reduction or proteolytic digestion. An example of this approach is the analysis of mAbs light and heavy chains, providing insight into posttranslational modifications of the individual chains. Figure 1 provides an overview of antibody sample preparation and various digestion options prior to intact mass analysis.
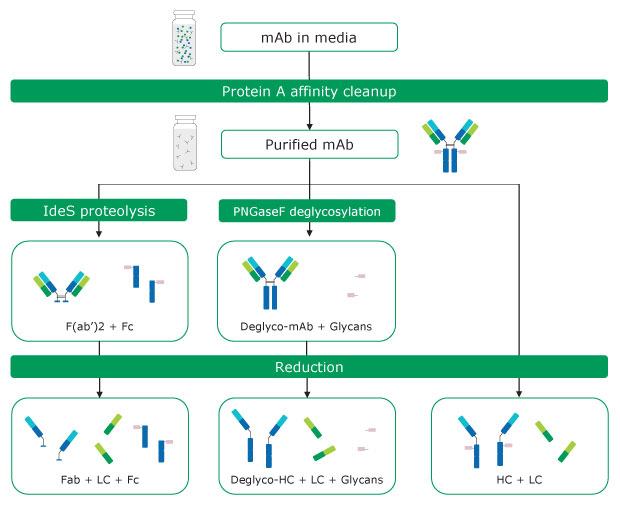
Figure 1.Antibody sample preparation
This report describes the application of both non-reducing and reducing SEC-MS workflows for the generation of intact mass and middle-up data of adalimumab. It includes the correlation of de-charged masses of respective heavy and light chains with calculated theoretical intact masses.
In all experiments, a recombinant human antibody standard, SILuTM Lite SigmaMAb (#MSQC4), was utilized as a reference and assay control sample. Throughout the text this is referred to simply as SigmaMAb although several different SigmaMAb standards are also available commercially.
General Procedures - Sample and Reference Preparation and System Setup for Antibody Purification and SEC-MS
Antibody Purification Procedure
The target antibody purification was performed on cell culture supernatants using immobilized protein A resin in a 96-well format. The suggested minimum working mAb titer is 100 μg/mL.
All procedures were conducted using both reference and assay control samples of SigmaMAb with a molecular mass of ~150 kDa. The reference sample consisted of the pure antibody reconstituted in water; it is used for the system suitability tests and instrument check. The assay control sample contained the antibody spiked into or delivered as a mixture with cell culture media or spent media (cell broth including nutrients etc.); it goes through the entire workflow and functions as a control sample.
In detail, the high-throughput purification of mAbs from cell culture media using protein A resin was performed as follows:
- System/Workflow Suitability
As part of the workflow suitability, an assay control of SigmaMAb in media was purified along with the samples. Reference sample (SigmaMAb) is prepared as follows: [SA1]- Reconstitute each vial of MSQC4 in 1.0 mL water to obtain a solution with an antibody concentration of 1 mg/mL.
- Prepare assay control (spent media sample) by spiking SigmaMAb in EX-CELL® CHOZN® Platform Medium, or equivalent, to obtain a final concentration of 100-500 µg/mL.
- Preparation of equilibration and elution buffers
- Prepare equilibration buffer (20 mM citrate, 150 mM NaCl, pH 7) by dissolving 5.82 g trisodium citrate dihydrate, 0.04 g citric acid, and 8.77 g sodium chloride in 1 L water. Adjust pH of resulting solution to 7 using 1 M NaOH or HCl as needed; subsequently filter solution using a 0.2 μm filter.
- Prepare elution buffer (25 mM citrate, pH 3) by dissolving 4.8 g citric acid in 1 L water. Adjust pH of resulting solution to 3 using 1 M NaOH or HCl as needed.
- Clarify samples
Centrifuge samples in tubes at maximum speed for five minutes and samples in plates at maximum speed for 60 minutes. - Protein A loading
- Add or remove water from top portion of settled protein A slurry to obtain a 50% protein A suspension.
- Mix slurry by constant pipette action and gentle shaking of reagent reservoir.
- Use a multichannel pipette to deliver 200 μL of protein A slurry to each well of a 96-well filter plate. Place protein A filter plate on a vacuum manifold. Catch any flow-through from the filter plate by placing the filter plate on top of a used collection plate.
- Protein A equilibration
- Add 200 μL of equilibration buffer to each well of protein A and apply vacuum to void wells of buffer.
- Repeat both steps twice.
- Washing bound mAb
- Place protein A filter plate on vacuum manifold with a waste collection plate inserted and the film cover removed.
- Apply vacuum to void wells of buffer media and transfer the filter plate onto a waste collection plate.
- Add 200 μL of equilibration buffer to wells and centrifuge plates at 3700 rpm for five minutes (this step helps in clearing the sample film on sides of filter plate wells).
- Add 200 μL of equilibration buffer to wells and apply vacuum to remove buffer.
- Repeat once more for a total of three washes.
- Eluting bound mAb
- Place protein A filter plate on a new collection plate and secure with a rubber band.
- Add 100 μL of elution buffer to each well, incubate filter plate on orbital shaker at 170 rpm for five minutes.
- Centrifuge plates at 3700 rpm for five minutes.
- Repeat addition of elution buffer, incubation on orbital shaker, and centrifugation for a total of three elution steps (300 μL of total elution volume).
Typical antibody recovery using this procedure is 60%.
Antibody Reduction Procedure
Disulfide (S-S) bond reduction was performed as follows:
- Prepare a 1 M dithiothreitol (DTT) solution by dissolving 154.25 mg DTT in 1 mL water.
- Prepare a 1 M ammonium bicarbonate (ABC) solution by dissolving 79.06 mg ABC in 1 mL water.
- Combine equal volumes of 1 M ABC and 1 M DTT to prepare the reduction solution.
- Transfer aliquots of 50 μL of each sample, system suitability reference, and control to autosampler vials.
- Reduce by addition of 5 μL 0.5 M ABC/0.5 M DTT solution.
- Incubate for one hour at room temperature or 30 min at 37 °C.
Note: Reduction is performed under non-denaturing conditions, where the inter-chain disulfide bonds (which are more susceptible to reduction) will break and produce the light and heavy chains, while the intra-chain disulfide bonds within each individual domain remain intact.
Alternatively, reduction of samples can be performed by using this protocol:
- Prepare a 100 mM solution of tris(2-carboxyethyl) phosphine (TCEP) in 6 M aqueous guanidine hydrochloride by dissolving 2.87 g of TCEP and 57.32 g of guanidine hydrochloride in 100 mL of water (if less solution is needed, scale down accordingly).
- Combine 30 μL of the resulting solution with 10 μL of sample.
- Incubate for two hours at 37 °C.
Note: Reduction performed under denaturing conditions, where both the inter-chain and intra-chain disulfide bonds (which are more susceptible to reduction) will break.
Instrument Calibration
The Waters QToF Xevo G2XS mass spectrometer was calibrated in a mass range of 500 – 6000 m/z with a 20 μL/min infusion of 0.4 mg/mL of cesium iodide in water. Alternatively, calibration can be performed with a 20 μL/min infusion of 0.4 mg/mL of polyalanine in water, prior to running the samples.
System Suitability
To evaluate the performance of the entire workflow, an assay control (SigmaMAb in media) was prepared and analyzed along with the samples. SigmaMAb reference was also tested to ensure system suitability (see section above).
In addition, reduced SigmaMAb antibody reference (formulated at 2 mg/mL and further diluted to 1 mg/mL) was analyzed alongside the samples to determine system suitability of the SEC-MS platform.
SEC-MS System Setup and MS Data Analysis
SEC-MS System Setup
The essential settings of the UHPLC-PDA chromatography system and the qToF mass spectrometer applied in the analysis of both reduced and non-reduced antibodies are listed in Tables 1 and 2 below.
MS Data Analysis
Data were processed using the MaxEnt1 module within the MassLynx 4.1 software to generate and analyze deconvoluted (zero charged) mass spectra. In general, a summed spectrum was created from the corresponding total ion chromatogram (TIC) of the eluted intact mAb, heavy chain (HC), or light chain (LC). The summed m/z spectrum was then processed by the MaxEnt1 algorithm. Detailed parameters are listed in Table 3.
For glycoform analysis, data were processed using UNIFI software from Waters. Glycoforms were matched by the software and HC glycoform glycans are listed in the respective section below. Glycoform relative abundance data was tabulated based on peak intensities of the coeluting glycoform species. A deconvolution filter setting employing a base peak intensity of 2% was utilized to preclude noise incorporation, and an output resolution setting of 5 Da was used.
Adalimumab SEC-MS Intact and Middle Up Analysis Results
Intact Mass Analysis of Non-Reduced Adalimumab
The analysis objective was to perform non-reduced SEC-MS intact mass analysis on all submitted samples to verify the molecular weight of adalimumab.
Protein A purification of samples of adalimumab and SigmaMAb antibody control was performed as described in the previous section. Intact SigmaMAb was used to determine system suitability. All mAb samples were solubilized in 100 μL H2O to obtain a final concentration of 1 mg/mL. Subsequently, samples were analyzed in their non-reduced form via SEC-MS.
System Suitability Test Results
SigmaMAb reference sample (10 μL) was injected on the SEC-MS system. Figure 2 illustrates the photodiode array (280 nm) and TIC (total ion current) traces of the non-reduced antibody, while Figure 3 displays the deconvoluted mass spectrum of the SigmaMAb reference. The observed intact mAb glycoforms matched the common glycoform masses of MSQC4, as listed in Table 4. The measured discrepancies between the observed masses and the theoretical values for four glycoforms were within 0.005% mass error or less.
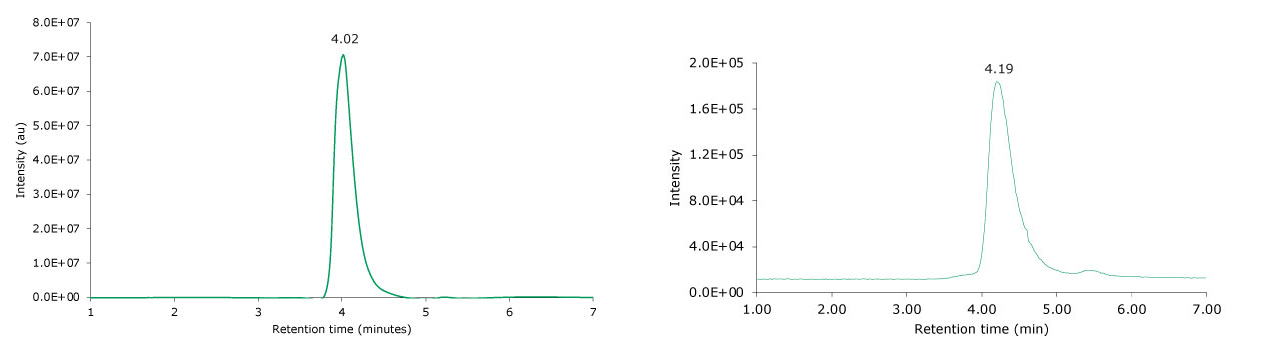
Figure 2.Photodiode array (280 nm, left) and TIC traces (right) of non-reduced SigmaMAb reference.
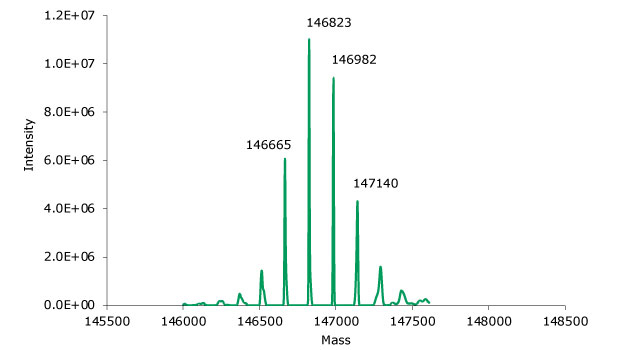
Figure 3.Deconvoluted mass spectrum of non-reduced SigmaMAb reference.
G0F: GlcNAc2Man3GlcNAc2Fuc
G1F: GalGlcNAc2Man3GlcNAc2Fuc
G2F: Gal2GlcNAc2Man3GlcNAc2Fuc
*Masses based on NIST Physical Reference Data
Non-Reduced Sample Results
The monoclonal antibody samples were analyzed in their non-reduced form using SEC-MS. The corresponding photodiode array (280 nm) traces, TICs, MS spectra, and the deconvoluted MS spectra of adalimumab are shown in Figures 4 and 5. The observed masses of the non-reduced mAb correlate well with the calculated theoretical masses for all submitted samples, as shown in Table 5, and the observed mass error was found to be 0.010% or less.
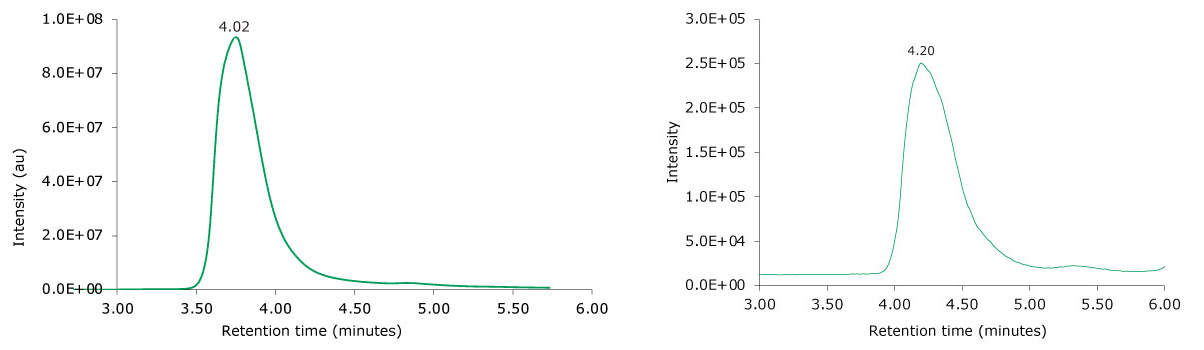
Figure 4.Photodiode array (280 nm, left) and TIC traces (right) of non-reduced adalimumab.
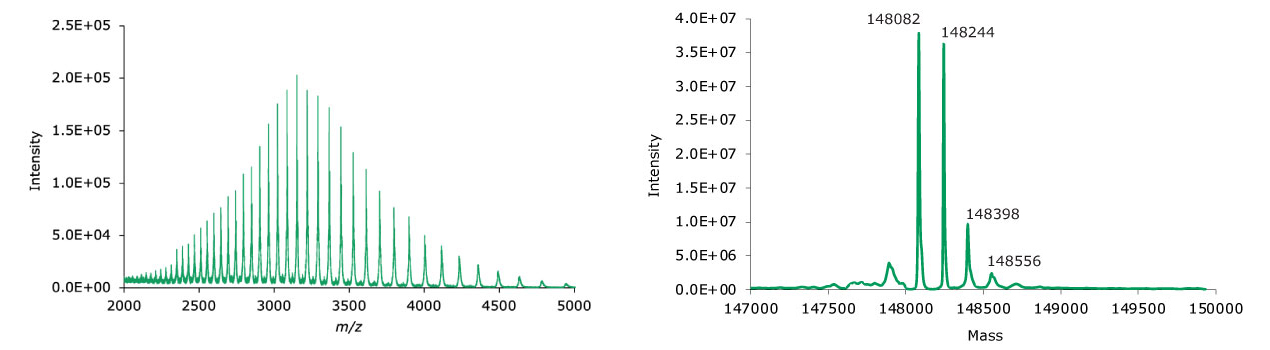
Figure 5.MS data for non-reduced adalimumab. Left: summed spectrum; right: deconvoluted spectrum.
G0F: GlcNAc2Man3GlcNAc2Fuc
G1F: GalGlcNAc2Man3GlcNAc2Fuc
G2F: Gal2GlcNAc2Man3GlcNAc2Fuc
*Masses based on NIST Physical Reference Data
Intact Mass Analysis of Reduced Adalimumab
The objective of the intact mass analysis was to verify the molecular weight of adalimumab light and heavy chains by applying an SEC-MS middle-up approach.
Protein A purification of samples of cell culture supernatants containing expressed adalimumab and SigmaMAb assay control was performed as described in the previous section. Reduced MSQC4 reference was used to determine system suitability. All mAb samples were solubilized in 100 μL H2O for a final concentration of 1 mg/mL. Subsequently, samples were analyzed in their reduced form via SEC-MS.
System Suitability Test Results
10 μL of a reduced SigmaMAb reference sample was injected on the SEC MS system. The observed heavy chain mAb glycoforms matched the expected glycoform masses of SigmaMAb, as listed in Table 6. Discrepancies between the observed masses and the theoretical values for all three glycoforms were within 0.003% mass error or less. Figure 6 illustrates the UV chromatogram (280 nm) trace of the SigmaMAb reference, while Figure 7 shows the summed and deconvoluted mass spectra of the reduced SigmaMAb reference light and heavy chain glycoforms.
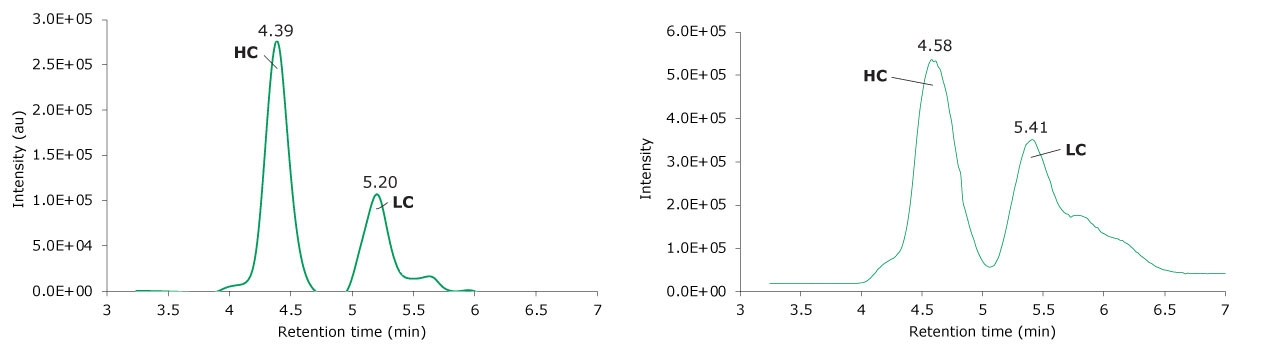
Figure 6.Photodiode array (280 nm, left) and TIC trace (right) of reduced SigmaMAb reference.
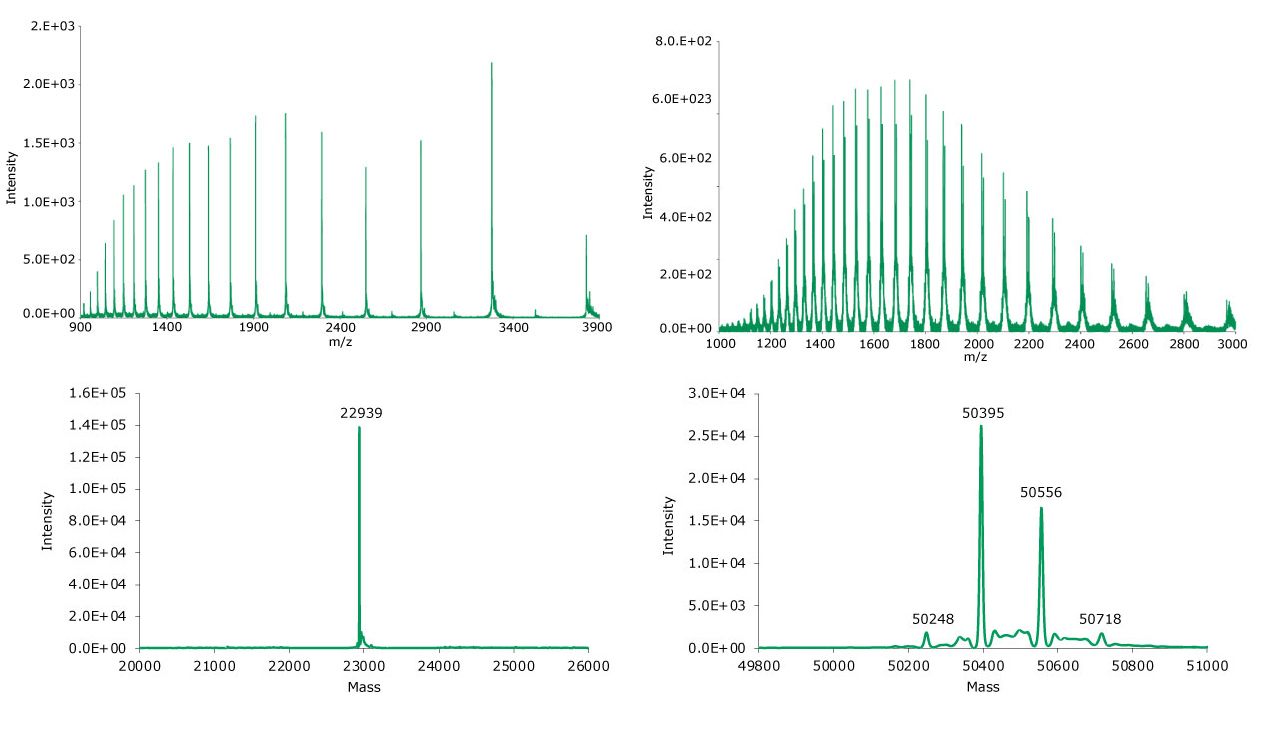
Figure 7.Summed (top) and deconvoluted (bottom) mass spectra of the light and heavy chains (left and right, respectively) of reduced SigmaMAb reference.
G0F: GlcNAc2Man3GlcNAc2Fuc
G1F: GalGlcNAc2Man3GlcNAc2Fuc
G2F: Gal2GlcNAc2Man3GlcNAc2Fuc
*Masses based on NIST Physical Reference Data
Reduced Sample Results
The reduced adalimumab samples were analyzed using SEC-MS. The corresponding photodiode array (280 nm) traces, TICs, MS spectra, and the summed and deconvoluted MS spectra of the samples are shown in individual Figures 8 and 9. The measured masses of reduced forms correlated well with the calculated masses as shown in Table 7 (mass error 0.005% or less). Deconvoluted masses of reduced light and heavy chains correlate well with the expected calculated masses for all submitted samples.
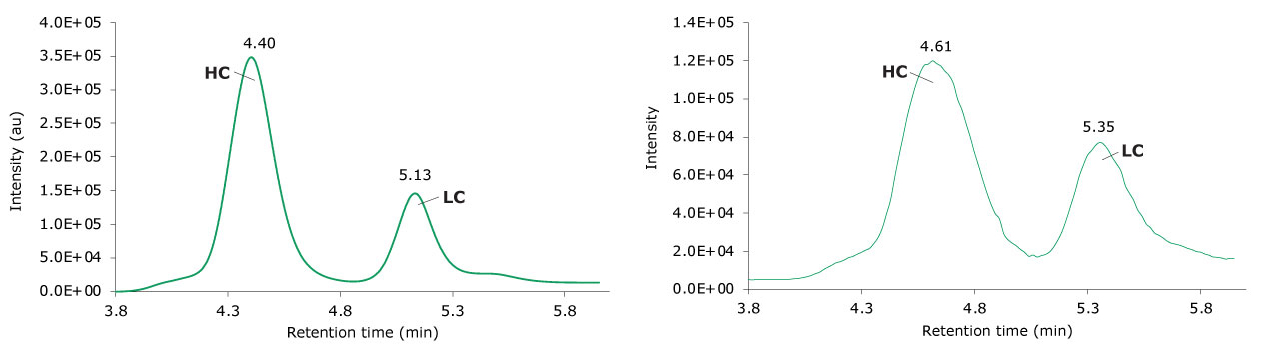
Figure 8.Photodiode array (280 nm, left) and TIC trace (right) of reduced adalimumab.
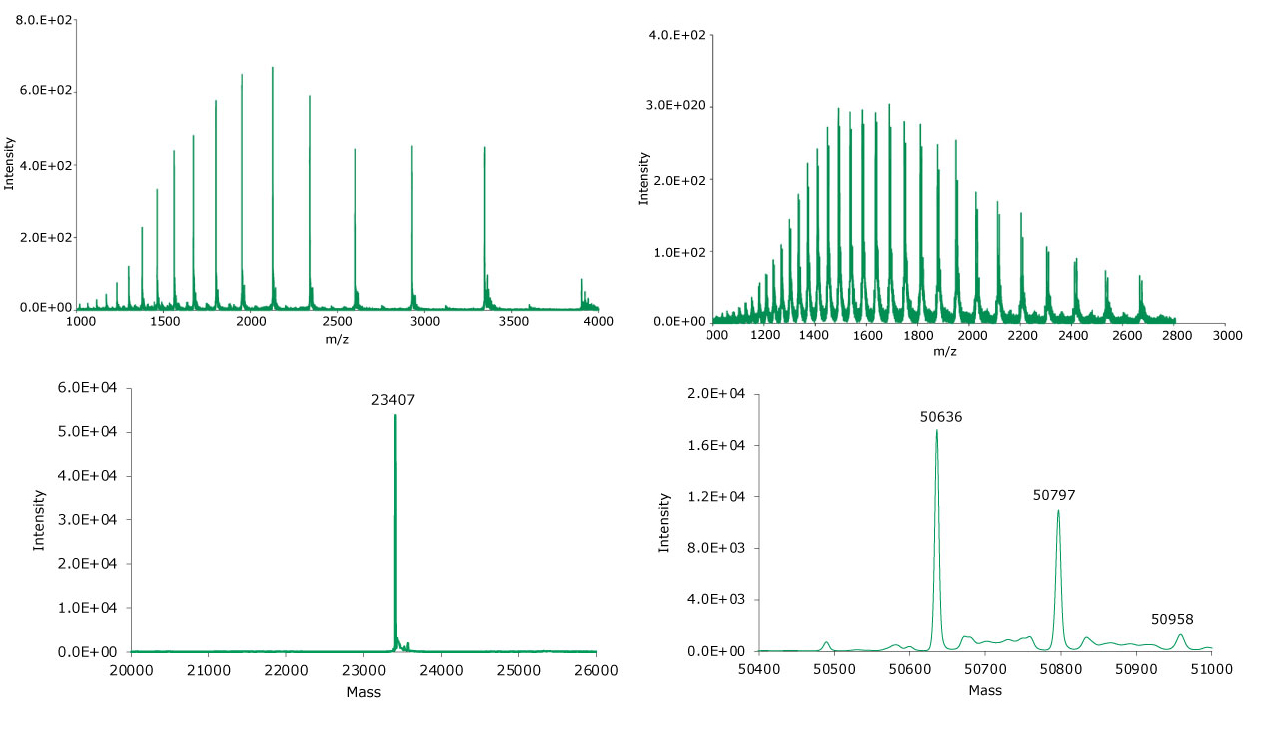
Figure 9.Summed (top) and deconvoluted (bottom) mass spectra of the light and heavy chains (left and right, respectively) of reduced adalimumab.
G0F: GlcNAc2Man3GlcNAc2Fuc
G1F: GalGlcNAc2Man3GlcNAc2Fuc
G2F: Gal2GlcNAc2Man3GlcNAc2Fuc
*Masses based on NIST Physical Reference Data
Conclusion
A workflow for the SEC-MS intact and middle-up mass analysis of reduced and non-reduced monoclonal antibodies was developed, using adalimumab as a model mAb and SILuTMLite SigmaMAb as a reference and assay control sample.
The workflow was comprised of an antibody purification process using immobilized protein A, an optional mAb reduction procedure, a mass spectrometer calibration method, and a system suitability test utilizing a recombinant human monoclonal antibody reference. In addition, a generic SEC method suitable for sample separation and analysis of both reduced and non-reduced mAbs was established.
Results for non-reduced SigmaMAb reference revealed measured discrepancies between the observed masses and the theoretical values for four glycoforms of 0.005% mass error or less. For adalimumab, the measured masses of the non-reduced mAb showed a strong agreement with the expected calculated masses for all submitted samples, and the observed mass error was observed to be 0.010% or less.
Analysis of reduced SigmaMAb reference sample revealed that the observed heavy chain mAb glycoforms matched the expected glycoform masses of the antibody. The discrepancies between the observed and the theoretical values for three glycoforms were all within 0.003% mass error or less. Similarly, the measured masses of reduced adalimumab correlated well with the calculated masses (mass error 0.005% or less).
The experimental data demonstrated that the workflow can be used for either reduced or non-reduced monoclonal antibody sample analysis, with accurate results allowing for an unambiguous identification of various glycoforms.