Quantitative PCR and Digital PCR Detection Methods
Introduction
Fluorescent dyes or probes are included in PCR mixes to monitor the change in DNA amplicon concentration as the reaction proceeds. Several popular detection methods, that are well established for the indirect measurement of template in qPCR, are discussed in this chapter. In addition, Dual-labeled Probes and some DNA-binding dyes have been shown to work well in digital PCR (dPCR).
Section Overview
Use the table of contents on the right to navigate to other sections of A Technical Guide to PCR Technologies.
dsDNA-Binding Dyes
Double-stranded DNA (dsDNA) binding dyes function as intercalating and/or minor groove binding agents and emit detectable fluorescence when bound to dsDNA but have a very low background when free in solution. Therefore, fluorescent signal intensity increases proportionately to the quantity of amplicon present. Double-stranded DNA-binding dyes are popular because they are a low cost detection option and do not require additional design considerations.
SYBR® Green I Dye
This dye is the most popular dsDNA-binding dye and has a long history of use in molecular biology. When free in solution, with only single-stranded DNA (ssDNA) present, SYBR Green I dye emits a signal of low intensity (Figure 5.1, Table 5.1). As the PCR progresses and the quantity of dsDNA increases, more dye binds to the amplicons and hence, signal intensity increases (see the animation on the following web page sigma.com/sybr-animation). However, since the dye binds to all amplified products indiscriminately, artefacts such as those resulting from primer dimers or nonspecific binding of the primers also contribute to the overall fluorescence. This can make it difficult to obtain accurate quantification, especially at low template concentrations. However, a post-PCR melt curve analysis can help determine reaction specificity1 (Figure 5.2).
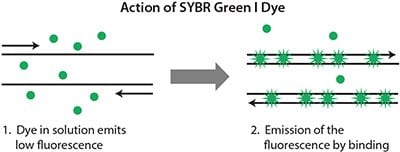
Figure 5.1.SYBR Green I dye cycles between an unbound (denatured) and a bound (annealing through extension) state as the reaction progresses and signal intensity increases as the quantity of amplicons increase.
aWhen complexed to dsDNA
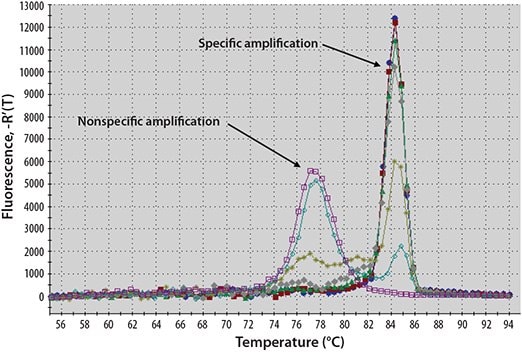
Figure 5.2.Example Melt Curve Analysis. dsDNA-binding dyes bind reversibly, so fluorescence intensity decreases as the reaction temperature increases above the melting temperature (Tm). In conjunction with controls, this type of analysis enables the detection of nonspecific products which melt at different temperatures than the specific product.
Probes
In all applications of qPCR, the amplification reaction is driven by specific forward and reverse primers. However, unlike assays relying on detection using dsDNA-binding dyes, probe detection systems do not include a free dye but rather a third oligonucleotide (and sometimes a fourth) conjugated to a reporter dye and/or a quencher moiety.
Dual-Labeled Probes
Dual-Labeled Probes (also known as hydrolysis or alternatively TaqMan® probes) are used in the 5’ nuclease assay2,3, which is the most popular probe detection chemistry (Figure 5.3; also see the animation of the following web page: sigma.com/probe-animation). A Dual-Labeled Probe is a single-stranded oligonucleotide that is labeled with a reporter dye and a quencher moiety. The reporter is located at the 5’ end and the quencher at the 3’ end. The quencher absorbs the natural fluorescence emission of the reporter, usually by Forster-type energy transfer, more commonly referred to as fluorescence resonance energy transfer (FRET). After amplification from the forward primer, the Taq DNA polymerase encounters the probe. The 5’ exonuclease activity inherent in the Taq DNA polymerase then separates the 5’ reporter from the 3’ quencher (Table 5.2, Figure 5.3), which provides a fluorescent signal that is proportional to the amplicon yield.
The hydrolysis probe assay is specific and accurate for quantification of low copy number targets. Specificity can be improved even further by the inclusion of modified nucleotides such as Locked Nucleic Acid in the probe (as described below). Locked Nucleic Acid modified probes are particularly useful for differentiating between single nucleotide polymorphisms (SNPs) or other similar sequences. Dual-Labeled Probes require careful design (PCR/qPCR/dPCR Assay Design) and are generally more expensive than dsDNA-binding dyes. Furthermore, while amplification of nonspecific products may remain undetected, side reactions may cause the overall reaction to be less efficient and so assays containing probes may still benefit from optimization (Sample Purification and Quality Assessment).
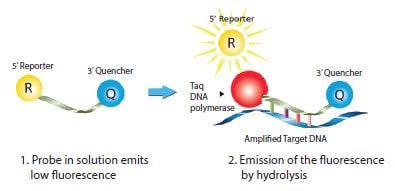
Figure 5.3.Mechanism of Dual-Labeled Probes. Taq DNA polymerase extends the primer situated on the same strand as the probe until it reaches the probe position. The inherent exonuclease activity hydrolyzes the probe from 5’ to 3’, which releases the reporter dye into solution and thereby causes an increase in fluorescence. The measured fluorescence signal is directly proportional to the amount of target DNA.
Most real-time PCR thermal cyclers have multiple detection channels enabling flexibility in the choice of probe labels. It is critical to select the reporter dyes that are compatible with the detection channels for the instrument and ensure that the correct filters and calibration are in place. When multiplexing, reporter combinations are required that are as distinct as possible from each other to minimize optical cross talk. Typical reporters include: FAM, HEX, TxRd (Sulforhodamine 101-X) and Cyanine 5. Under identical conditions, it is usual to observe differences in emission intensity from different reporters. For this reason, it is advisable to analyze the data from each reporter combination independently (using different threshold settings as appropriate for the probe emission).
Molecular Beacons
Molecular Beacons (also known as hybridization probes) are single-stranded probes that are held in a hairpin-loop conformation (20–25 nucleotides) by complementary stem sequences (4–6 nucleotides) at each terminus4 (Figure 5.4). The hairpin-loop is complementary to the template and the hydrogen-bonded stem sequences allow the 3’ quencher to suppress the fluorescence of the 5’ reporter (Table 5.3) when the probe is free in solution.
Figure 5.4.Mechanism of Molecular Beacons. Molecular Beacons hybridize to their specific target sequence causing the hairpin-loop structure to open, separating the 5’ reporter from the 3’ quencher. As the quencher is no longer in proximity to the reporter, fluorescence emission takes place. Unlike Dual-Labeled Probes, the mechanism of detection of Molecular Beacons does not rely on degradation during the reaction. The measured fluorescence signal is directly proportional to the amount of target DNA.
LightCycler® Probes
A LightCycler Probe or FRET system (also known as dualhybridization probes) consists of a pair of single-stranded fluorescent-labeled oligonucleotides5,6 (Figure 5.5). Oligo Probe 1 is labeled at the 3’ end with a donor fluorophore dye and Oligo Probe 2 is labeled at the 5’ end with one of a few available acceptor fluorophore dyes (Table 5.4). The free 3’ hydroxyl group of Oligo Probe 2 must be blocked with a phosphate group to prevent DNA polymerase extension.
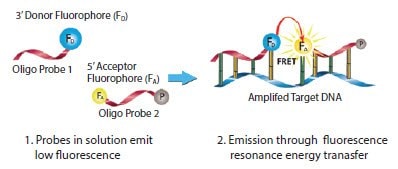
Figure 5.5.Mechanism of LightCycler FRET Probes. During the annealing step, the primers and both of the probes hybridize to their specific target regions, bringing the donor dye into close proximity to the acceptor dye (the probes are usually spaced 1 to 5 nucleotides apart). When the donor is excited by light from the realtime PCR instrument, energy is transferred by FRET from the donor to the acceptor. The emission wavelength for the dye on the acceptor probe is detected. The increase in fluorescence signal is directly proportional to the amount of target DNA.
Scorpions® Probes
Scorpions® probes are available in two forms; uni-probe and bi-probe. The uni-probe consists of a stem-loop structure, similar to a Molecular Beacon but this is attached to the forward primer with a PCR blocker located between the two oligo sections7 (the blocker prevents Taq DNA Polymerase from extending the primer (Figure 5.6A, Table 5.5).
The bi-probe structure is a duplex with the 5’ reporter, probe sequence, PCR blocker and forward primer on one strand and the 3’ quencher on the other strand; the quencher strand is duplexed with the reporter strand (Figure 5.6B).
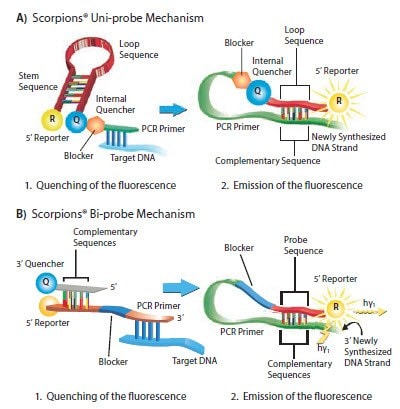
Figure 5.6.Mechanism of Scorpions® Probes. The uni-probe A) has all probe components on one strand, whereas the bi-probe B) has the probe components on two strands. Both formats of Scorpions® Probes include the forward PCR primer. This forward primer is extended to become part of the newly formed amplicon. During annealing/extension, the probe sequence in the Scorpions® hybridizes to the template, which separates the reporter from the quencher and thereby results in a fluorescent signal. As the tail of the Scorpions® and the amplicon are part of the same strand, the detection interaction is intramolecular and therefore more rapid than other probe detection systems. The template is typically chosen to be between 5 and 50 bases from the 3’ end of the Scorpions® primer. Both the uni-probe and bi-probe Scorpions® require a separate reverse primer.
Quenchers
Most probe detection systems require a quencher moiety. Some of those used in the original probes structures were acceptor fluorescent dyes, e.g., TAMRA, which work well with FAM but are not suitable for other dyes. As a reporter dye itself, TAMRA produces fluorescence and therefore can result in a poor signal-to-noise ratio. For this reason, dark quenchers which emit heat instead of light, such as Black Hole Quencher® (BHQ®) were developed by Biosearch Technologies. The choice of dark quenchers as popular alternatives to dye molecules provides quenching over a wide range of wavelengths and opens the possibility of multiplex reactions containing a larger number of target/probe combinations.
Onyx Quencher™ (OQ™) is a proprietary dark quencher from Sigma-Aldrich. Four derivative versions (OQA, OQB, OQC and OQD) are available. As shown in Table 5.6, the four Onyx Quenchers are compatible with a variety of popular reporter dyes.
The data presented in Figure 5.7 show amplification of an artificial template that was derived from a synthetic oligo for the optimization of a Schistosoma mansoni target assay8. Detection was using a FAM-labeled probe with a comparable dark quencher, CDQ (A) or OQA (B). From these data, it is evident that both the CDQ and the OQA have equivalent performance, with similar background fluorescence and similar Cq values for analyzed data from template of the same concentration.
In summary, OQ is equivalent in performance to CDQ and importantly, is available license, restriction and royalty free for any application. This makes the Onyx Quencher an excellent and cost effective choice for the development of commercial kits and reagents for molecular diagnostics that contain qPCR probes.
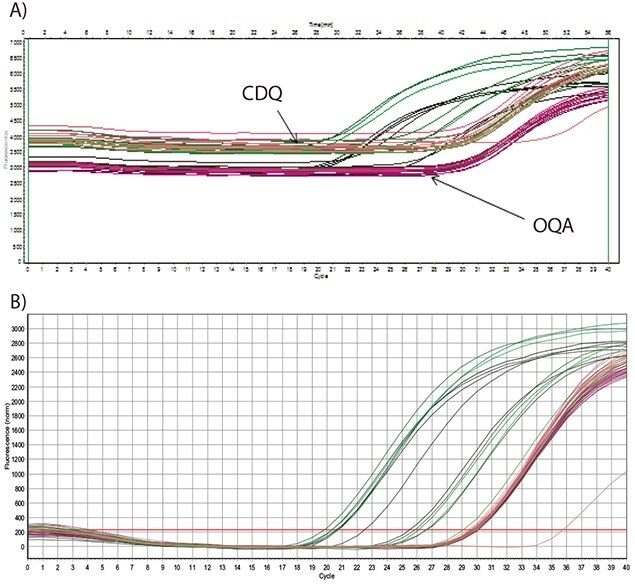
Figure 5.7.Dilutions of an artificial oligo were amplified using 250 nM primers and the amplicon detected with a Dual-Labeled Probe at 200 nM. A) The probe was labeled with FAM and quenched with CDQ or OQA (as indicated) and the raw fluorescence data are shown. B) The probe was labeled with FAM and quenched with CDQ or OQA and the baseline corrected data are shown. There are no significant differences between the performances of either probe.
Nucleic Acid Analogs
A number of modifications can be used to produce oligonucleotides with altered biochemical properties. These modifications generally confer a higher Tm which can be manipulated to provide improved specificity. This allows for assay design in regions of challenging sequence or when a single oligonucleotide is required to detect several sequences, e.g., all serotypes of a virus.
Locked Nucleic Acid
Locked Nucleic Acid is an RNA analog base (Figure 5.8) that yields enhanced sensitivity and specificity when incorporated into probes9. Probes with Locked Nucleic Acid bases have greater thermal stability and therefore hybridize more strongly to the template. Each Locked Nucleic Acid base may increase the Tm of the probe by up to 8 °C10, which makes Locked Nucleic Acid a powerful tool in SNP discrimination assays11 (a single base mismatch has a greater destabilizing effect on duplex formation than without Locked Nucleic Acid), multiplexing (allows for simpler Tm optimization) and problematic target sequences (Locked Nucleic Acid probes can be shorter, which allows them to be designed around problems such as AT- or GC-rich regions, repetitive sequences, or sequences with significant secondary structure). PCR Primers containing Locked Nucleic Acid have also been found to beneficial in applications such as SNP genotyping12.
Locked Nucleic Acid also provides protection against nuclease digestion making it suitable for in vivo use13.
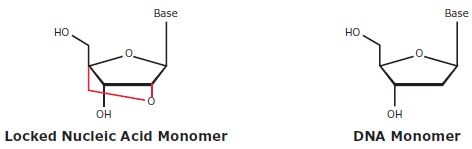
Figure 5.8.Comparison of Locked Nucleic Acid and DNA nucleotide structures. Locked Nucleic Acid differs from DNA in that Locked Nucleic Acid contains ribose with a methylene bridge between the 2’ oxygen and 4’ carbon, which ‘locks’ the ribose in the 3’ endo conformation, whereas DNA contains 2’-deoxyribose with no methylene bridge.
Summary
There are many considerations when selecting a detection method for a particular application. While SYBR Green I dyes and Dual-Labeled Probe assays are popular and generally work well, there are situations in which the other detection systems might be preferable. Table 5.7 can be used as guidance during early assay development.
Blank spaces indicate that the detection method is not recommended; X = good performance; and XX = better performance.
References
To continue reading please sign in or create an account.
Don't Have An Account?