LC-MS Analysis of a Semi Polar Metabolites QC Mix on an Ascentis® Express AQ-C18 Column
Abstract
To evaluate variability in LC-MS metabolomics data analysis, an additional quality control (QC) sample is recommended at the beginning of each analytical batch. The QC sample enables the detection of instrumental drift in LC-MS analysis, including fluctuations in signal intensity values, ion suppression effects, or retention time shifts of chromatographic peaks in the LC-MS data. A semi-polar QC mix composed of nine compounds can be employed to monitor drift and ion suppression phenomena. A chromatographic example method was established for this mixture using an Ascentis® Express AQ-C18 column, allowing a reliable separation of the semi-polar metabolites in a single run under highly aqueous mobile phase conditions.
Section Overview
Introduction
Metabolomics is defined as the profiling of small molecules derived from biochemical processes and pathways.1,2 Samples commonly analyzed include stool,3 serum/plasma,4 urine,5 cerebrospinal fluid,6 and saliva.7 The field is applied across various research areas, including microbiome studies,8 nutrition,9 disease studies,10 and agriculture,11 where metabolites are analyzed through two main approaches: targeted and untargeted metabolomics.12-15 Metabolite analysis is performed using two principal strategies: targeted and untargeted metabolomics. In targeted metabolomics, known specific chemical groups, including short-chain fatty acids,16 bile acids, lipids,17 and amino acids18 are analyzed. In contrast, untargeted metabolomics involves the comprehensive analysis of unknown chemical compounds present within a sample.
Liquid chromatography-mass spectrometry (LC-MS) is widely employed as the primary technique for metabolite profiling in metabolomics analyses.19,20 To monitor variability in LC-MS based metabolomics data, it is advisable to include an additional quality control (QC) sample at the beginning of each analytical batch, with repeated injections every 4-10 samples throughout the workflow.19,21,22 The QC sample is used to assess analytical drift, such as fluctuations in intensity values, ion suppression effects, or changes in the retention times of chromatographic peaks in the LC-MS data.
The Semi Polar Metabolites QC Mix (SBR00052) is a ready-to-use solution for mass spectrometry workflows. The QC mix enables effective monitoring of analytical drift and ion suppression effects. It contains nine components, including bile acids, aromatic carboxylic acids, and nucleosides (Table 3). The solution is supplied in a ready-to-use format with approximately 9:1 water to acetonitrile ratio and is packaged in amber vials fitted with silicone/PTFE liner crimp tops compatible with most LC-MS autosamplers. The mixture should be stored at refrigerated (2-8 °C) and does not require thawing prior to use.
Experimental
The example LC-MS analysis was performed using an Ascentis® Express AQ-C18 column, which is resistant to dewetting and allows the use of highly aqueous mobile phases up to 100% water (Tabe 1). Mass spectrometric detection was conducted in both positive and negative electrospray ionization (ESI) modes.
Results and Discussion
The mass spectrometry (MS) electrospray ionization positive (ESI+) and negative (ESI-) base peak chromatograms (BPC) for the Semi Polar Metabolites QC Mix are presented in Figures 1 and 2. Distinct peaks labeled 1 to 9 were observed, corresponding to the semi-polar metabolites in the mix (refer to the lists of metabolites in Tables 3 and 4). A stable baseline was obtained, indicating good sensitivity and effective separation, providing a clear profile of the metabolites present in the QC mix, suitable for subsequent analytical assessments.
Method A - ESI(+)
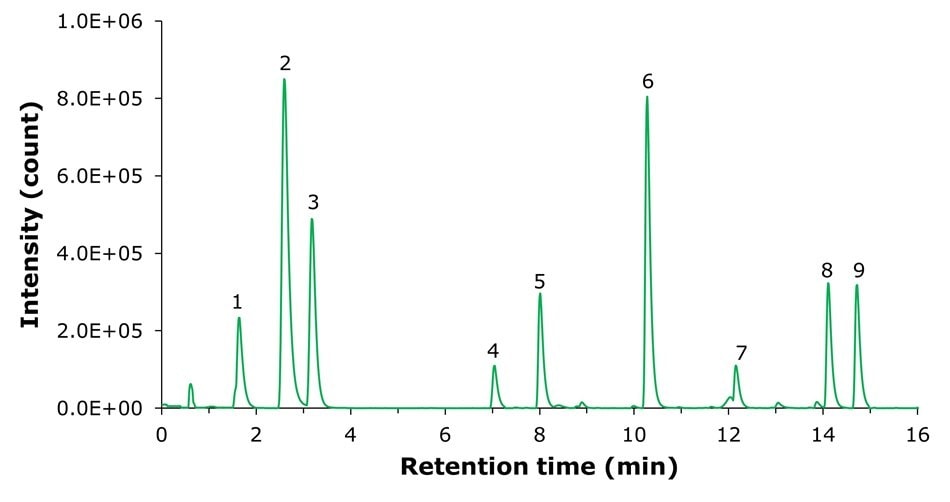
Figure 2.MS ESI(+) Method A base peak chromatogram (BPC) of the Semi Polar Metabolites QC Mix.
Method B - ESI(-)
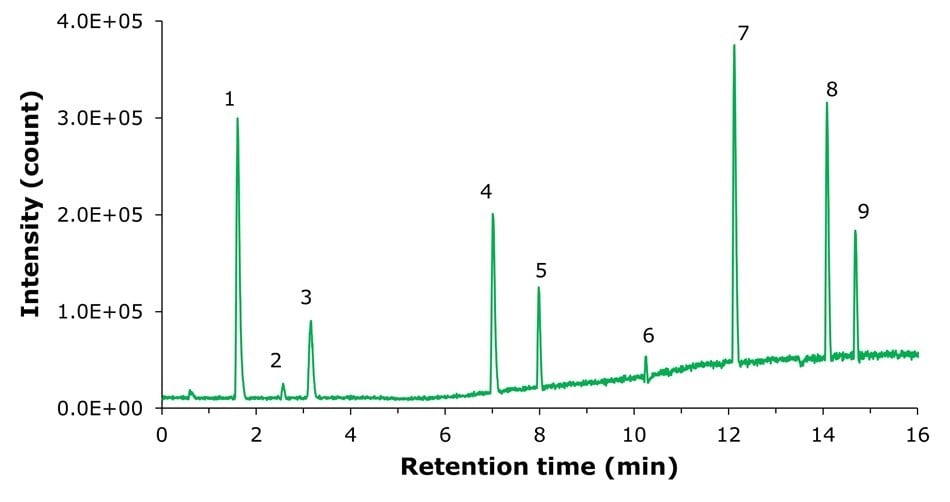
Figure 3.MS ESI(-) Method B base peak chromatogram (BPC) of the Semi Polar Metabolites QC Mix.
Conclusion
The Semi Polar Metabolites QC Mix is a specialized solution developed for LC-mass spectroscopy applications in metabolomics research. It enables a robust LC-MS methodology for the 9 included compounds to effectively monitor and evaluate drift and ion suppression phenomena, making it a valuable tool to ensure consistent results across metabolite profiling studies. The mix is suitable for both ESI(+) and ESI(-) applications and it comes conveniently packaged as a ready-to-use solution.
Related Products
HPLC Column
Solvents and Reagents
Quality Control Samples
References
To continue reading please sign in or create an account.
Don't Have An Account?