HILIC Analysis of Twelve Nucleosides on an Ascentis® Express ZIC®-cHILIC Column
Section Overview
Introduction
Nucleosides are classified as glycosylamines and consist of a nucleobase covalently linked to a five-carbon sugar moiety (pentose). Various nucleoside analogues have been developed and are employed in clinical practice as therapeutic agents, specifically in the treatment of cancer and viral infections. Structural modifications in these analogues have been introduced to enhance their efficacy and specificity toward pathways involved in cellular proliferation and viral replication.1,2
This application focusses on the qualitative analysis of a standard mixture containing twelve nucleosides (Figure 1) by HPLC with UV detection. The separation was performed on a HILIC column with a zwitterionic phase (phosphorylcholine) on superficially porous particles (SPP), the Ascentis® Express ZIC®-cHILIC.
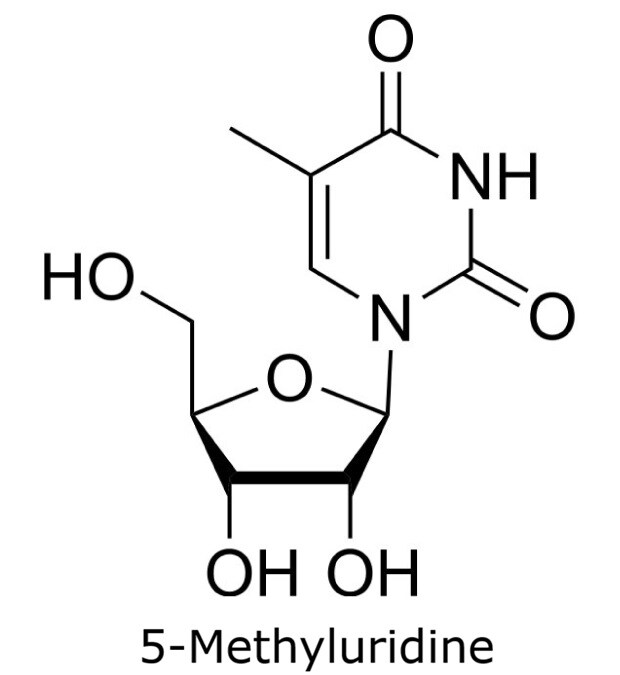
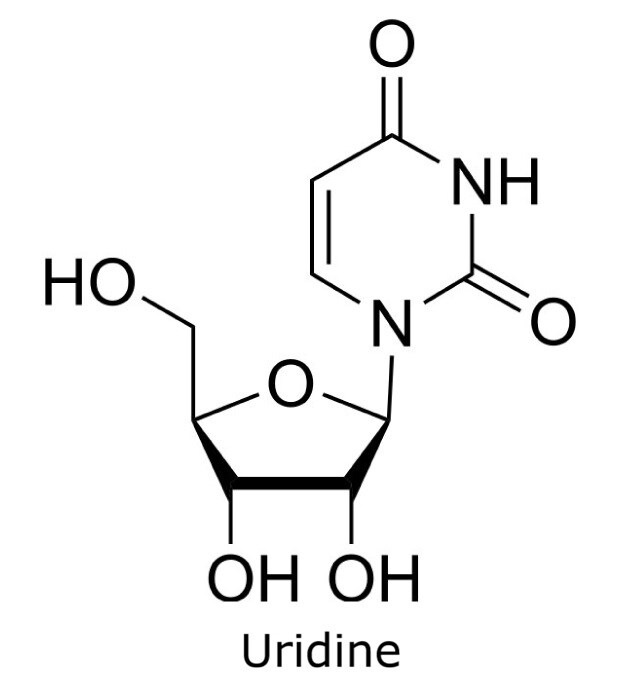
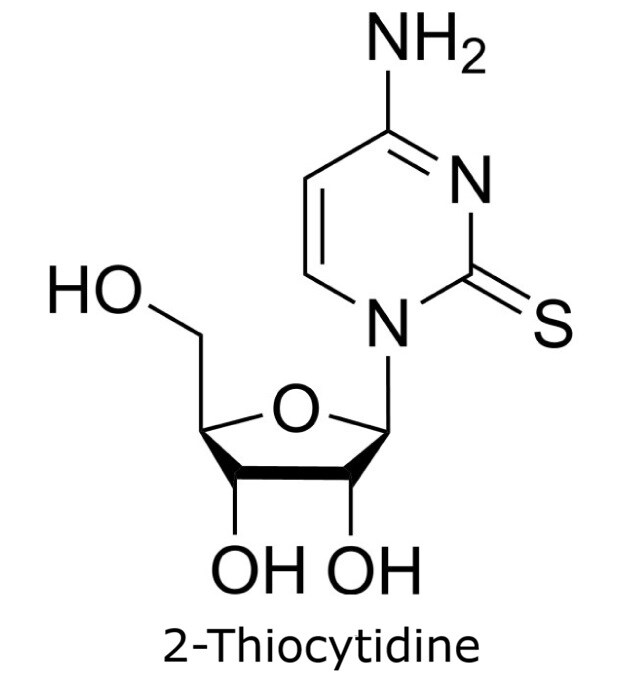
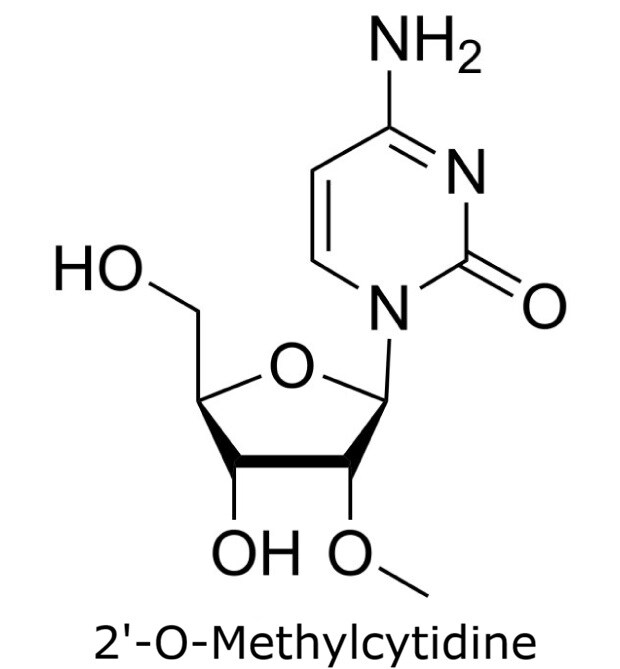
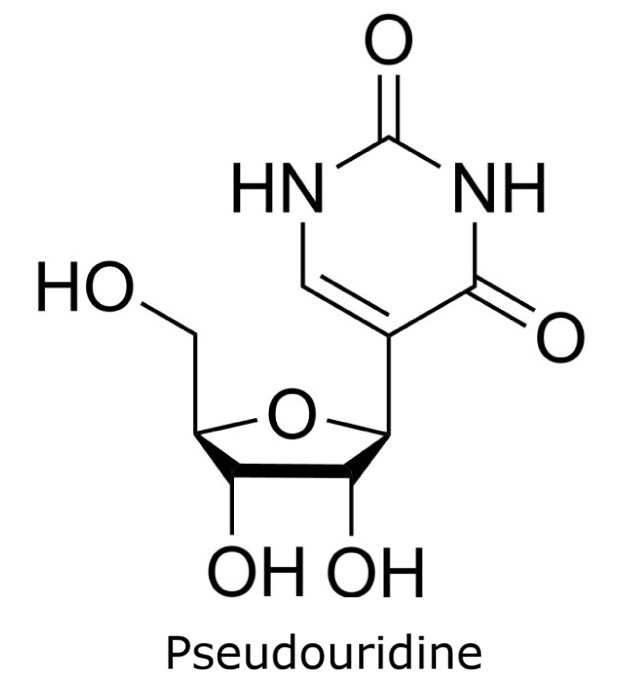
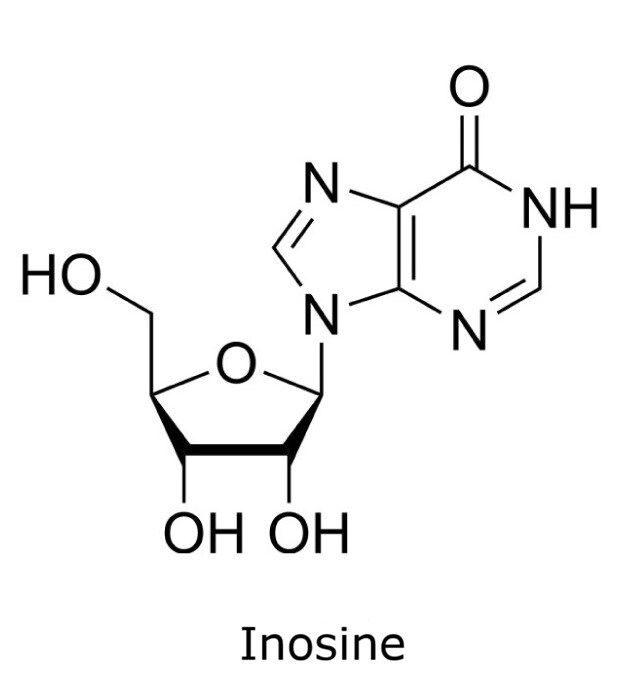
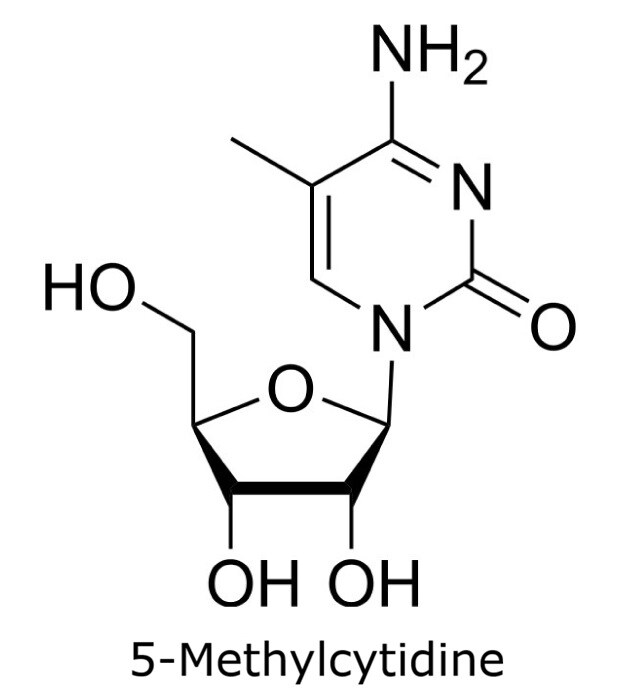
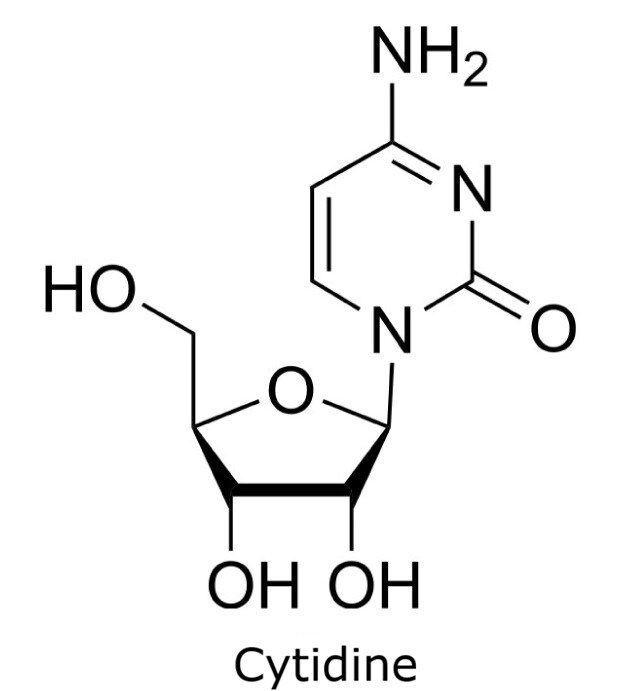
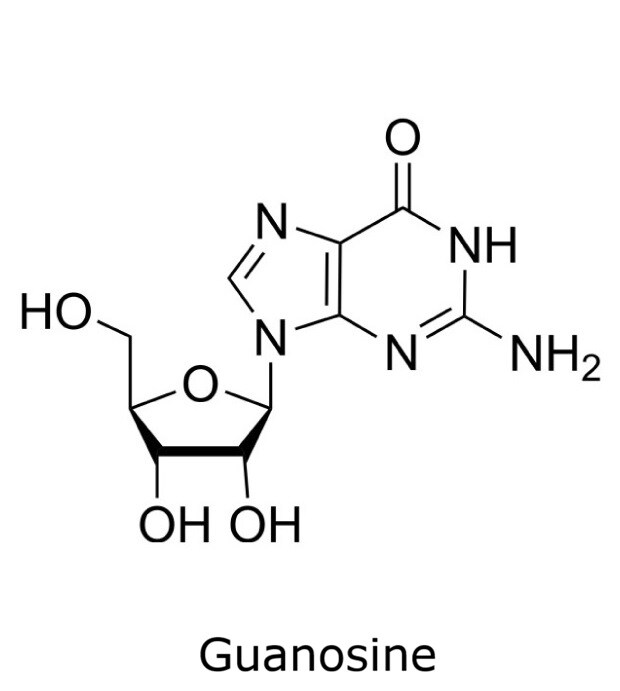
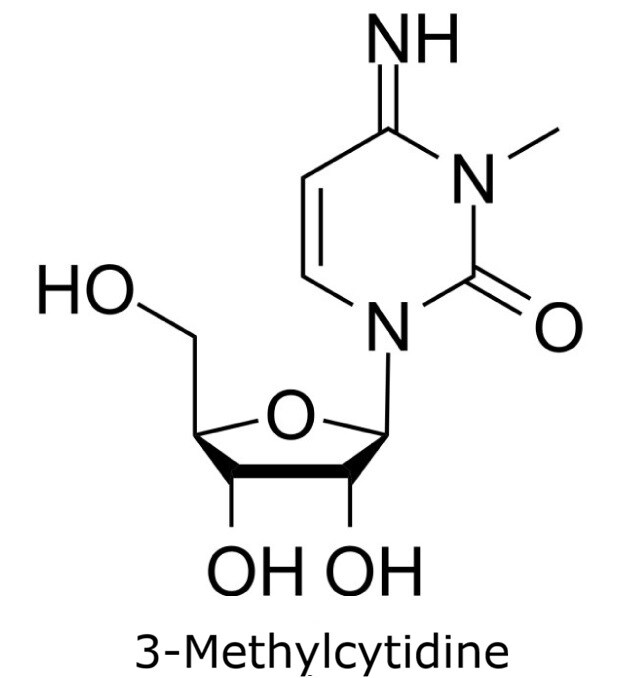
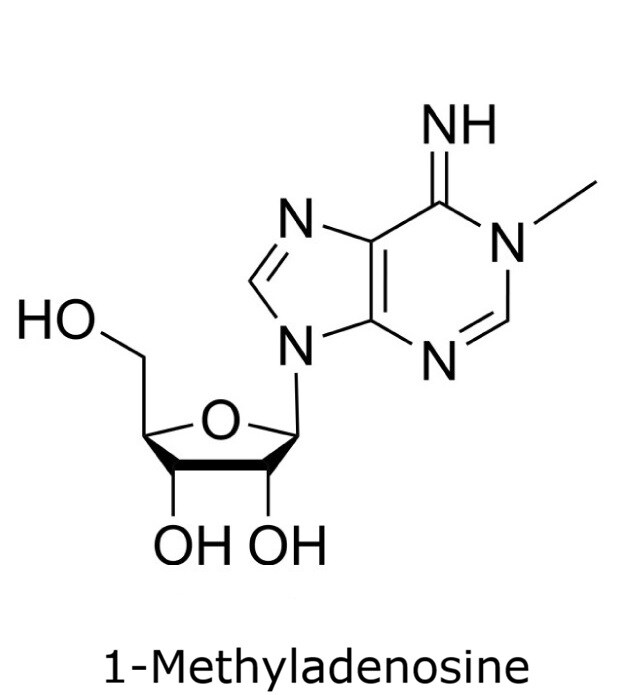
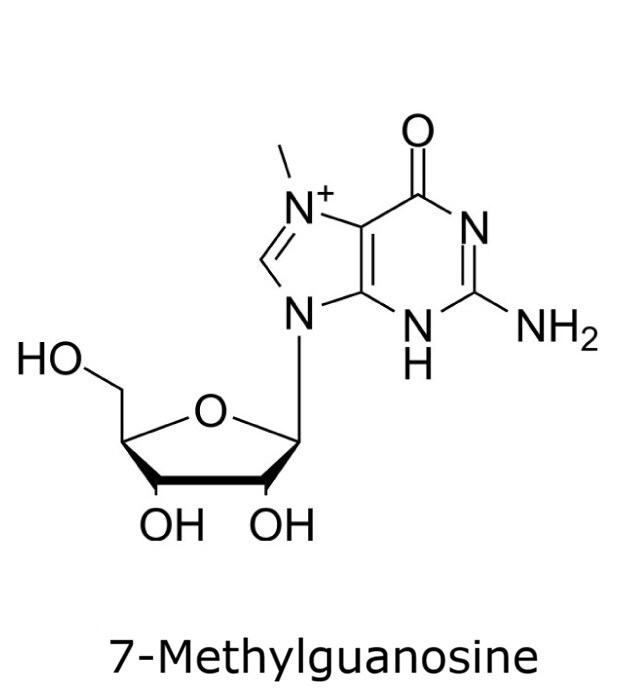
Figure 1. Compounds in Nucleoside Test Mix (47310-U).
Experimental
The Nucleoside Test Mix (47310-U; matrix: 1% sodium formate in water) was diluted at a ratio of 1:10 with acetonitrile, to prepare a standard solution (Table 1) with a matrix composition similar to the initial mobile phase condition of the used HILIC method, which consisted of buffer and acetonitrile in a 3:97 ratio (Table 2). The qualitative analysis of the twelve nucleosides at varying concentrations in the standard mixture was carried out using UV detection on an Ascentis® Express 160 Å ZIC®-cHILIC 2.7 µm SPP column (150 x 2.1 mm I.D.).
Results
The qualitative analysis of the standard mixture containing twelve nucleosides at varying concentrations was carried out by U/HPLC-UV on an Ascentis®Express 160 Å ZIC®-cHILIC, 2.7 µm SPP column (150 x 2.1 mm I.D.). The separation (Figure 2) showed good chromatographic performance, providing baseline separation with sharp and symmetrical peaks (Table 3).
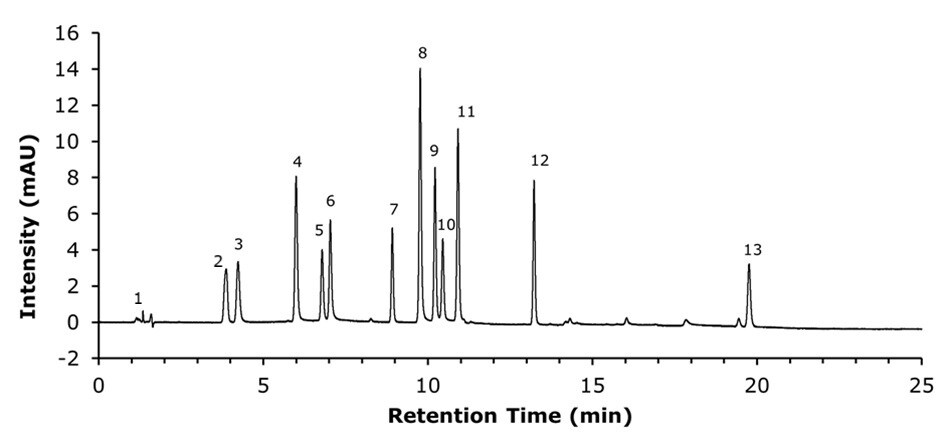
Figure 2.U/HPLC-UV chromatogram obtained for the standard mixture containing twelve nucleosides using an Ascentis® Express 160 Å ZIC®-cHILIC column (2.7 µm, 150 x 2.1 mm).
Conclusion
The developed simple HPLC-UV HILIC method enabled the determination of twelve nucleosides with baseline separation and good peak symmetry on an Ascentis® Express 160 Å ZIC®-cHILIC 2.7 µm SPP column (150 x 2.1 mm). The zwitterionic phosphorylcholine phase on the superficially porous particles (SPP) provided excellent retention and efficiency, enabling optimized HILIC separations.
Reagents, Solvents, and Standard
References
To continue reading please sign in or create an account.
Don't Have An Account?