LC-MS Analysis of a Polar Metabolites QC Mix on a ZIC® HILIC Column
Abstract
In a metabolomics analysis workflow, inclusion of a quality control (QC) sample at the beginning of each LC-MS batch is recommended to detect instrumental drift, including fluctuations in signal intensity, ion suppression, and retention time shifts. The ready-to-use Polar Metabolites QC Mix, containing eight compounds, is applied for effective monitoring of these effects. A hydrophilic interaction liquid chromatographic (HILIC) method example was developed using a SeQuant® ZIC®-cHILIC column, enabling analysis of the Polar Metabolite QC mix within a 25-minute run.
Section Overview
Introduction
Metabolomics is defined as the profiling of small molecules derived from biochemical processes and pathways,1,2 primarily characterized from samples such as stool,3 serum/plasma,4 urine,5 cerebrospinal fluid,6 and saliva.7 The field spans various research areas including microbiome studies,8 nutrition,9 disease study,10 and agriculture11. Metabolite analysis is generally performed through two main approaches, targeted and untargeted metabolomics.12-15 Targeted metabolomics focuses on analyzing specific groups of known metabolites, such as short-chain fatty acids,16 bile acids, lipids,17 and amino acids.18 In contrast, untargeted metabolomics examines all unknown chemical compounds present within a sample.
Liquid chromatography-mass spectrometry (LC-MS) is widely employed as the primary technique for metabolite profiling in metabolomics analysis.19,20 To detect variations in LC-MS metabolomics data, inclusion of an additional quality control (QC) sample at the start of each analytical batch is recommended, followed by repeated injections after every 4-10 samples within the workflow.19,21,22 Such a QC sample helps in identification of drifting effects, such as fluctuations in intensity values, ion suppression, or changes in the retention times of peaks. The ready-to-use Polar Metabolites QC Mix (SBR00055), suitable for LC-MS analysis, allows users to effectively monitor drifting and ion suppression phenomena. The QC mix contains eight compounds comprising polar metabolites like amino acids, vitamins, and nucleosides (Table 3). The mixture is prepared in ~9:1 acetonitrile / 10 mM ammonium formate in water and is therefore suitable for direct injection. The solution is packaged in a crimp-top amber vial with a silicone/PTFE liner to ensure compatibility with most LC-MS autosamplers. It is recommended to store the mixture at a temperature of 2-8 °C.
Experimental
The example LC-MS analysis was performed using a SeQuant® ZIC®-cHILIC column under HILIC conditions (Tabe 1). Mass spectrometric detection was conducted in both positive and negative electrospray (ESI) modes.
Results and Discussion
The mass spectrometry (MS) electrospray ionization positive [ESI(+)] and negative [ESI(-)] base peak chromatograms (BPC) for the Polar Metabolites QC Mix are shown in Figures 1 and 2. Distinct peaks labeled 1 to 8 were observed and were assigned to the corresponding polar metabolites in the mix (refer to the metabolites list in Tables 3 and 4). A stable baseline was observed, indicating good sensitivity and effective chromatographic separation, thereby providing a clear profile of the metabolites present in the QC mix, essential for subsequent analytical assessments.
Method A - ESI(+)
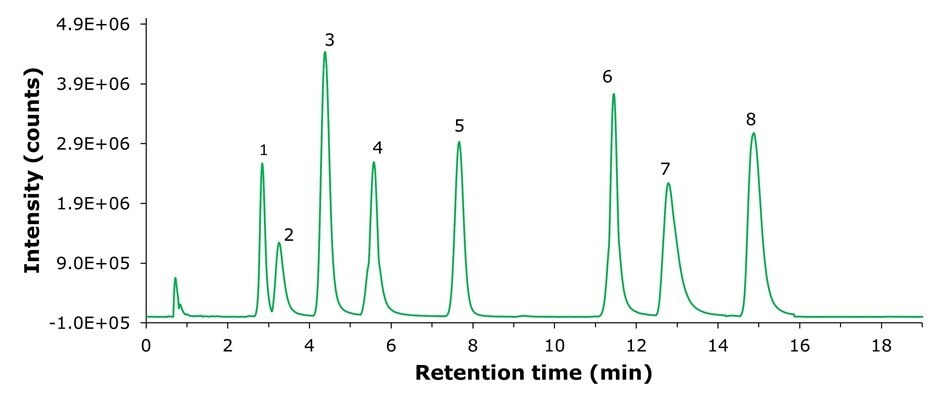
Figure 1.MS ESI(+) base peak chromatogram (BPC) of the Polar Metabolites QC Mix.
Method B - ESI(-)
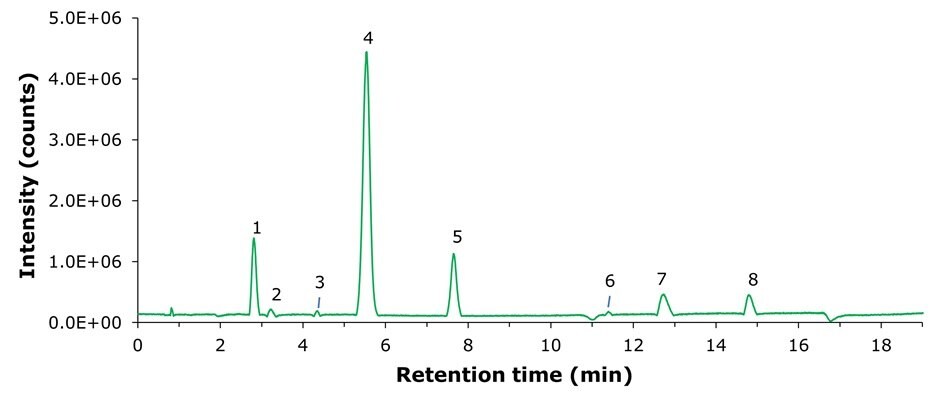
Figure 2.MS ESI(-) base peak chromatogram (BPC) of the Polar Metabolites QC Mix.
Conclusion
The Polar Metabolites QC Mix is presented as a dedicated solution for LC-MS applications in metabolomics research. It enables a robust LC-MS methodology for the eight included compounds, allowing effective monitoring and evaluation of drift phenomena and supporting consistent results across metabolite profiling studies. The Mix is designed to be applicable and suitable for both ESI(+) and ESI(-) applications and comes conveniently packaged as ready-to-use solution.
Related Products
HPLC Column
Solvents and Reagents
Quality Control Samples
References
如要继续阅读,请登录或创建帐户。
暂无帐户?