How To Automate DNA Extraction with the GenElute™-E Single Spin Blood DNA 96 Kit on the AAW™ Automated Assay Workstation
This automated DNA extraction protocol describes how to automate DNA extraction from blood samples using the GenElute™-E Single Spin Blood DNA 96 Kit on the AAW™ workstation. Lyse and extract 96 blood samples with high throughput and consistency through the power of automation.

Intro To Automating DNA Extraction
DNA extraction is a long, multi-step process that can be particularly time-consuming for large numbers of samples. This article provides a guide to automate lysis and extraction of DNA from blood samples in a high throughput (96-well) manner. GenElute™-E DNA extraction kits use negative chromatography technology to produce high-quality, full length genomic DNA with less shearing and fewer small, unwanted DNA pieces.
The GenElute™-E user guide protocol has been adapted into Python code, tested, and made available through the Millipore® Protocol Library for the AAW™ workstation. Once downloaded to the instrument, it can be used to perform all liquid handling, heating, and shaking steps, with two brief pauses for plate seal application and removal.
Our verified protocol allows up to 96 separate samples of blood to be lysed and extracted in 2 hours. The resulting 96 wells of isolated DNA can be quantified by spectrophotometer or fluorometer or used immediately for downstream assays such as PCR or sequencing.
Automation and GenElute™-E DNA Extraction Materials
View labware, hardware, and reagents for this automated DNA extraction protocol.
Labware
These are required for all assays. Quantities needed include one 50 µL tip rack, four 200 µL tip racks, and one or two deep well plates (available from Opentrons®, Prod. No. 999-00103).
Microcentrifuge tubes and the conical tubes are optional.
Hardware
There are two options for the AAW™ workstation, the “Assay Ready” version includes certain accessories with purchase whereas the other one does not. Additional accessories are available for purchase separately.
The tube rack is optional.
Reagents
Protocol Uploading and AAW™ Workstation Setup
New users should refer to the AAW™ user manual for detailed instructions on loading protocols to your AAW™ workstation, however for simplicity we have included instructions here for guidance.
- To load any program onto the AAW™ workstation, users should first download the Opentrons® application on a computer and establish access to the AAW™ robot through Wi-Fi, ethernet, or USB cable.
- Then, download the GenElute-E™ Blood DNA Extraction Protocol and custom labware files from the Millipore® Protocol Library. Import the protocol into the Opentrons® App ‘Protocols’ tab and upload the custom labware files under the ‘Labware’ tab using [Import] (upper right corner).
- Use the meatball (⋮) menu to the right of the protocol and select either “Send to Opentrons FLEX” OR “Start set-up” to load the protocol onto the AAW™ workstation.
- Once the protocol is loaded onto the AAW™ workstation, users can move to the on-device touchscreen or proceed to set-up using the computer application.
- Parameters: There are two versions of the GenElute-E™ Blood DNA Extraction Protocol – one for samples starting in tubes, and one for samples pre-aliquoted into a NEST 96-Well Deep Plate.
- Both protocols require users to select options for the following parameters:
- Dry Run: Yes or No (default is No). Selecting Yes allows the user to quickly test the protocol without consuming reagents or tips.
- Multi-Channel Mount: Left or Right (default is Left).
- Lysis Plate: GenElute™-E Round Well Plate or NEST Deep Well Plate
- For the Plate Protocol ONLY, users must also select an option for the following parameter:
- Number of Samples: 1-96 (default is 96)
- For the Tubes Protocol ONLY, users must select options for the following parameter:
- Sample Start: large tube, small tube (default = large tube). User should choose whichever type of labware the samples will be starting in: 15 mL conical tubes or 2 mL tubes.
- CSV File: users are required to input a CSV file (using the provided template) that lists sample volume and number of replicates.
- Both protocols require users to select options for the following parameters:
- Once finished with parameters, click [confirm values].
- If instruments (pipettes and gripper) are not already set up, clear the robot deck and follow on-screen directions to attach the AAW™ 8-Channel 1,000 µL Pipette, AAW™ 1-Channel 1,000 µL Pipette, and AAW™ Flex Gripper. Calibrate both pipettes and gripper as needed following the on-screen prompts.
- Install deck hardware by loading them into the indicated locations:
- Heater-Shaker. Remove the deck plate in D1 and install the Heater-Shaker module in its carrier. Ensure the power and USB cables are plugged in and the light comes on. Calibrate as required.
- Trash Bin. Remove the deck plate in A3, secure the carrier, and place the trash container inside.
- Labware position can be checked following the on-screen instructions and AAW™ User Guide. When complete, click [update offsets] and [confirm offsets].
- If labware position check for this protocol has already been recently accomplished, users may accept the previous offsets by only clicking [confirm offsets] without re-doing checks.
- Labware should be placed on deck, including plasticware and pipette tips. Once complete, deck should be loaded as shown below in Figures 1 and 2. Click [confirm placements] when finished.
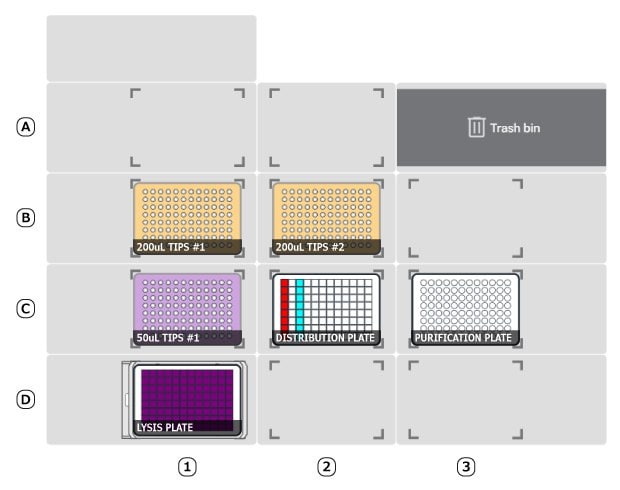
Figure 1.Deck setup for Plate protocol.
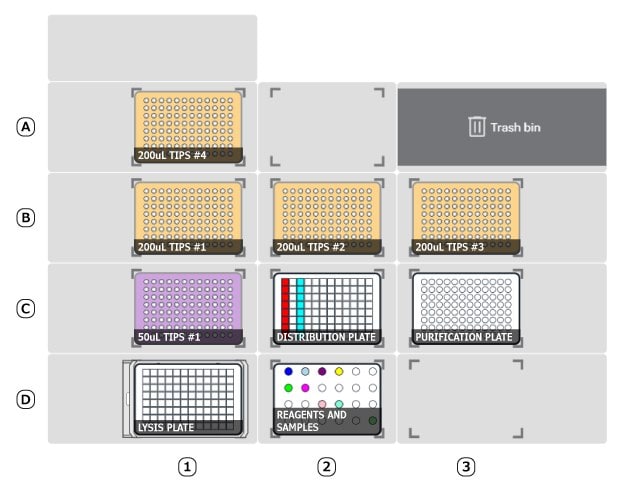
Figure 2.Deck setup for Tube protocol.
- Liquids should be added into the reagent plate as indicated on the touchscreen, with volumes based on the total number of samples multiplied by the number of replicates.
- Lysis Buffer + Protease master mix should be evenly distributed in Column 1.
For a full plate, thoroughly mix 5,760 µL of Blood Lysis Buffer with 1,152 µL of SmartLyse™ B Protease. Distribute 864 µL of mix into each well of Column 1. - Clearing Solution should be evenly distributed in Column 3.
For a full plate, distribute 135 µL of clearing solution into each well in Column 3. - Water (Tubes Protocol ONLY): If any samples are distributed at a volume of less than 60 µL, users will be prompted to include a water tube to make up the difference.
- Lysis Buffer + Protease master mix should be evenly distributed in Column 1.
Automated Blood DNA Extraction Kit Execution
Once all setup steps are complete and the user is certain all items are correctly placed for the protocol, close the door to the robot deck and click [Run] at the top of the touchscreen.
The AAW™ workstation will add lysis buffer, sample, and (if necessary) water, then pause for manual addition of the plate seal. After the plate is sealed, it will shake at 60 °C for 30 minutes, then at 80 °C for 10 minutes. The protocol will pause again for manual removal of the seal and movement of the plate off of the heater-shaker to slot D3. The protocol will then continue with the addition of clearing solution, brief shake to mix, and transfer from lysis plate to purification plate.
At the end of the protocol, users should place the purification plate on top of the storage plate and centrifuge according to the GenElute™-E user guide. Figure 3 shows how the manual workflow compares to the protocol for the automated blood DNA extraction kit.

Figure 3.Representative manual vs automated protocol timeline comparison.
Results: Automated DNA Extraction vs Manual DNA Extraction
This protocol was tested and verified with DNA extraction performed according to the GenElute™-E User Guide either manually or with the adapted protocol on the AAW™ machine. The assay was repeated three times on the AAW™ workstation and twice manually with a full plate of bovine blood samples (30 µL blood diluted with 30 µL water). Resulting DNA was quantified by spectrophotometer (Figures 4 and 5) and representative samples were run on an agarose gel (Figure 6).
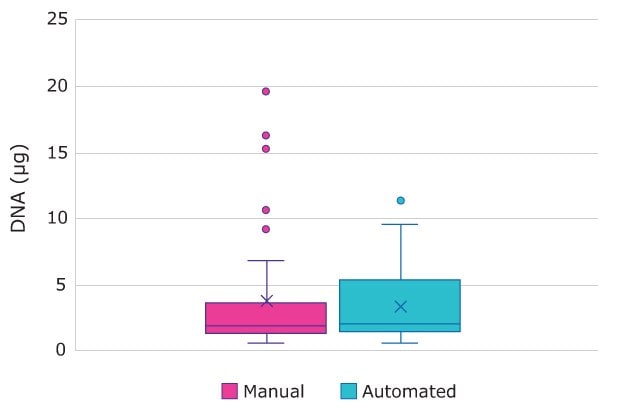
Figure 4.DNA Yield per 10 µL Input. Manual and automated assays produced similar distribution DNA yields, with an interquartile range of 1.31-3.47 µg for manual and 1.49-5.10 µg for automated assay per 10 µL of input blood.

Figure 5.DNA Quality. Manual and automated assays produced DNA of similar quality as determined by 260/280 ratios, with an interquartile range of 1.58-1.65 for manual and 1.73-1.92 for automated assay.

Figure 6.Representative Samples on Agarose Gel. Four representative samples each from manual and automated assays were run on a 1% agarose gel.
Summary
This article serves as a guide to automate DNA extraction from blood samples using the GenElute™-E Single Spin Blood DNA 96 Kit. It demonstrates both the consistency and reliability that comes with automation on the AAW™ workstation using verified protocols from the Millipore® Protocol library. Automated results performed similar to manual efforts, showing how this platform can reduce hands-on time and help enhance DNA extraction sample throughput.
Request more information on the AAW™ workstation below.
Tips and Tricks
- For first-time users, a dry run of the protocol is recommended to become familiar with the automated assay steps. Dry runs have shorter mixing and incubation steps and can be performed with no liquid or with water or buffers.
- Dilution with water is recommended for particularly viscous samples, to ensure proper mixing and lysis.
如要继续阅读,请登录或创建帐户。
暂无帐户?